Probes Newsletter 04/2019
Newsletter 04/2019
XCyting analysis of BCR-ABL1-like ALL/LBL...
BCR-ABL1-like B-lymphoblastic leukemia/lymphoma (B-ALL/LBL) are lymphoblastic neoplasms committed to the B-Cell lineage characterized by the lack of the BCR-ABL1 translocation while the gene profiles analyzed are very similar to that in ALL with BCR-ABL1. This new WHO entity is genetically based on chromosomal rearrangements of genes coding for growth factors, cytokines, downstream tyrosine kinases, and growth factor/cytokine receptors. Furthermore, several gene mutations are known.
To detect rearrangements of the frequently BCR-ABL1-like B-ALL associated gene CRLF2, MetaSystems Probes is further expanding its portfolio and adds two new probes for the detection of CRLF2 rearrangements. While XL CRLF2 BA detects CRLF2 gene rearrangements, the sister probe XL P2RY8 del detects deletions between the CRLF2 locus and the proximal PR2Y8 gene, a frequent rearrangement partner of CRLF2. Thus, the XL P2RY8 del design allows the detection of the CRLF2-P2RY8 fusion gene. For further information, please visit www.metasystems-probes.com.
XL CRLF2 BA
Clinical Applications: ALL
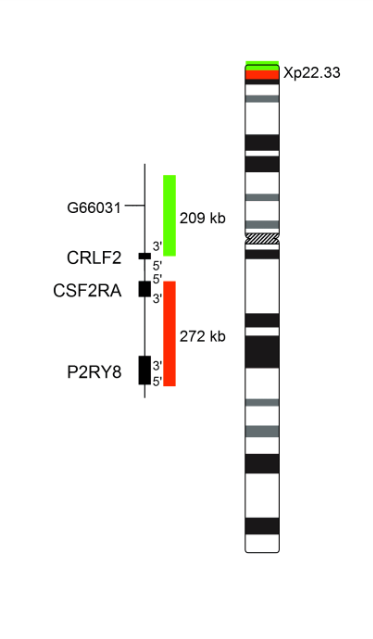
XL CRLF2 BA is designed as a break apart probe. The orange labeled probe hybridizes proximal to the breakpoint in the CRLF2 gene region at Xp22.33 and Yp11.32, the green labeled probe hybridizes distal to the breakpoint at Xp22.33 and Yp11.32.
Download fact sheet (PDF) for further information.
Acute lymphoblastic leukemia (ALL) is the most common malignancy in children (prevalence of approximately 1:1500). Furthermore, children with Down syndrome have a 10- to 20-fold increased risk of developing acute leukemia (B-cell precursor-ALL and acute myeloid leukemia, particularly acute megakaryoblastic leukemia). B-Cell dependent BCR-ABL1-like ALL, also known as Philadelphia chromosome (Ph)-like ALL, is a high-risk subset with a gene expression profile that shares significant overlap with that of Ph-positive (Ph+) ALL. In contrast to Ph+ ALL, defined by the presence of the BCR-ABL1 fusion, BCR-ABL1-like cases contain a variety of genomic alterations activating kinase and cytokine receptor signaling. In 2017, the WHO recognized BCR-ABL1-like ALL as new entity. Besides a variety of further functions, cytokine receptor-like factor 2 (CRLF2) plays an important role in cytokine induced B-Cell proliferation and development. Chromosomal rearrangements resulting in the overexpression of CRLF2 can be found in 50% of BCR-ABL1-like ALL cases. The CRLF2 gene is located in the pseudoautosomal region 1 (PAR1) of the X and the Y chromosome, and several translocations of the type t(X;14) or t(Y;14) as well as interstitial deletions have been described. Three genetic key mechanisms regarding CRLF2 and ALL are known. First, the CRLF2 gene is placed under control of the IGH enhancer. t(X;14) or t(Y;14) translocations are the genetic basis for this aberration. Second, fusion of CRLF2 to CSF2RA, a further PAR1 gene, has also been described. Third, cryptic interstitial deletions of the PAR1 region rearrange the CRLF2 gene juxtapose the initial non-coding exon of P2RY8 (purinergic receptor P2Y8). The resulting P2RY8-CRLF2 fusion being under the control of the P2RY8 promoter is strongly transcribed in lymphoid cells. CRLF2 rearrangements result in increased protein levels, which initiate significantly enhanced JAK/STAT (Janus kinase/signal transducer and activator of transcription) signaling, whereby disproportionate JAK and subsequent STAT5 activation induces strongly enhanced B-cell activation and proliferation.
XL CRLF2 BA detects chromosomal aberrations resulting from CRLF2 gene rearrangements (deletions or translocations). XL P2RY8 del can be used as an additional specific tool to detect the possible presence of the P2RY8-CRLF2 fusion gene.
XL P2RY8 del
Clinical Applications: ALL
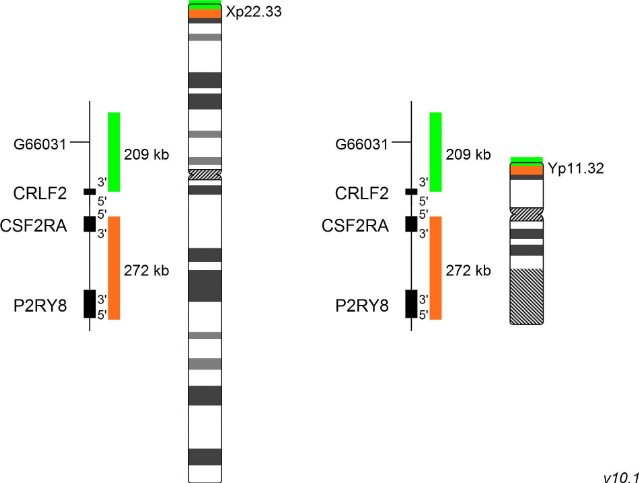
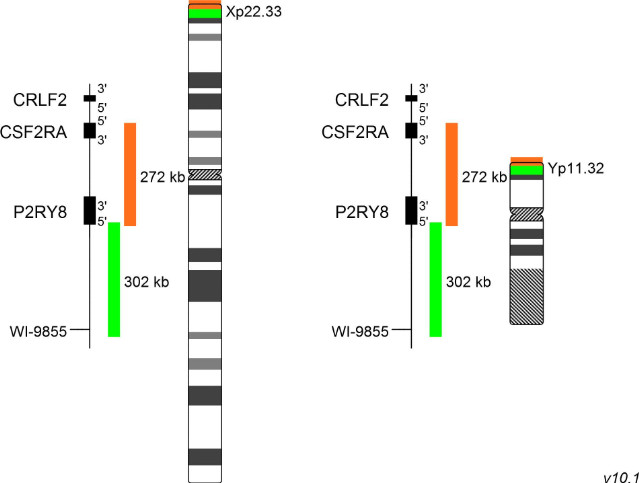
XL P2RY8 del detects deletions in the short arm of chromosome X and Y at Xp22.33 and Yp11.32, respectively. The orange labeled probe spans P2RY8 and extends distally, the green labeled probe covers the 5´end of P2RY8 and extends proximally.