XL Iso(17q)
Deletion/Isochromosome Probe
- Order Number
- D-5048-100-OG
- Package Size
- 100 µl (10 Tests)
- Chromosome
- 1717
- Regulatory Status
- IVDD
IVDR Certification
MetaSystems Probes has already certified a wide range of FISH probes, according to IVDR.
This product remains IVDD-certified until further notice.
Discover all IVDR-certified products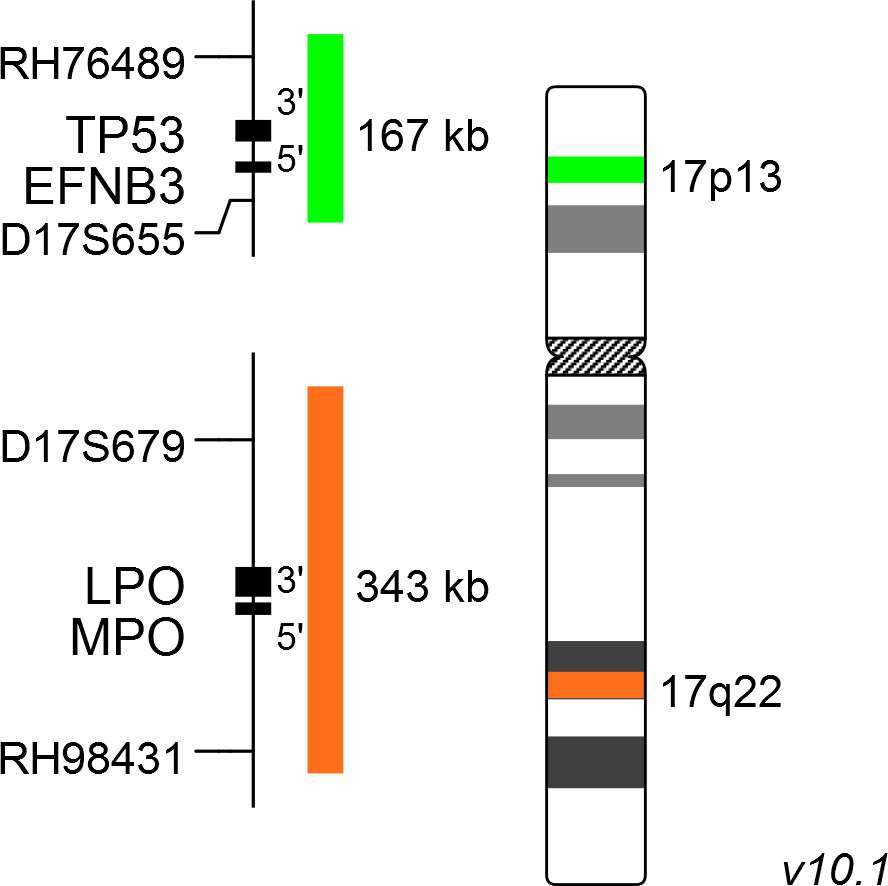
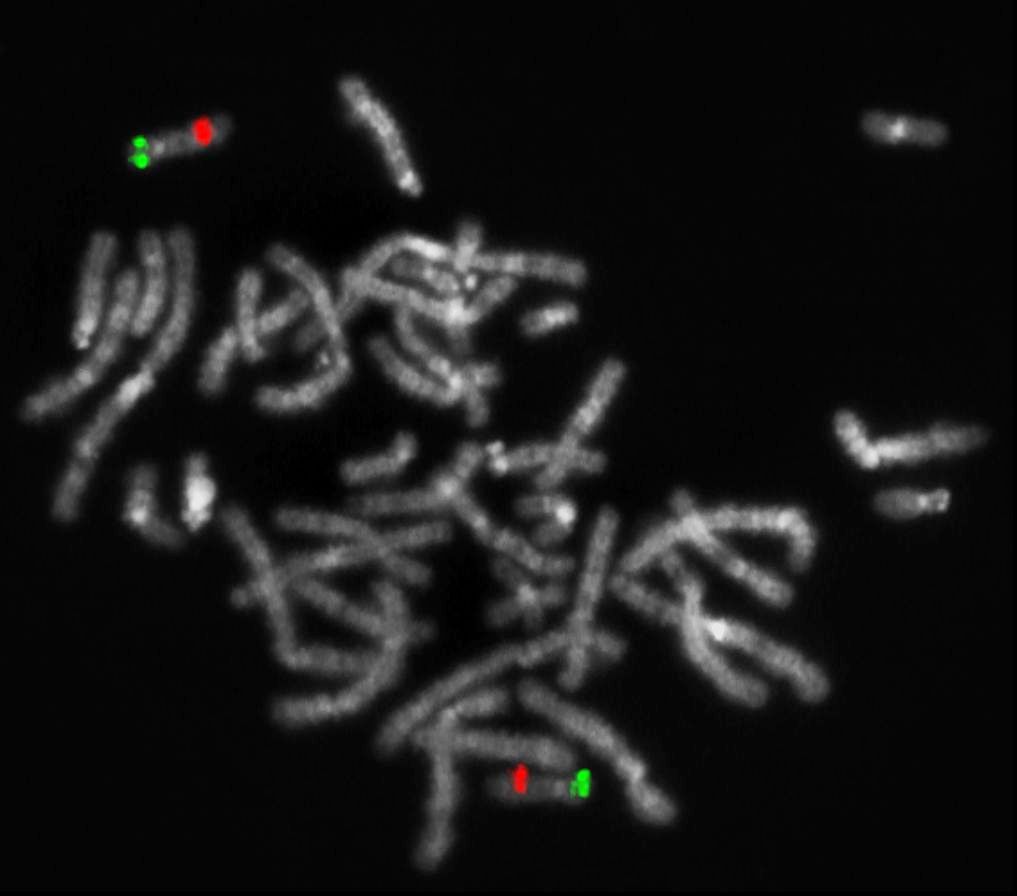
XL Iso(17q) consists of a green-labeled probe hybridizing to the TP53 gene region at 17p13 and an orange-labeled probe hybridizing to the MPO gene region at 17q22.
Probe maps for selected products have been updated. These updates ensure a consistent presentation of all gaps larger than 10 kb including adjustments to markers, genes, and related elements. This update does not affect the device characteristics or product composition. Please refer to the list to find out which products now include updated probe maps.
Probe map details are based on UCSC Genome Browser GRCh37/hg19, with map components not to scale.
An isochromosome of the long arm of chromosome 17, i(17q), is the most frequent genetic abnormality observed during the disease progression of Philadelphia chromosome positive (Ph+) chronic myeloid leukemia (CML). The breakpoints are located in the short arm of chromosome 17 within the Smith-Magenis critical region at 17p11. In neuroblastoma and other hematologic malignancies, amplification of 17q is a significant predictive factor for adverse outcome.
Isochromosome 17q, or i(17q), is ocurring in primitive neuroectodermal tumor/medulloblastoma (50%), chronic myeloid leukemia (CML), acute myeloid leukemia (AML), and myelodysplastic syndrome (MDS).
Clinical Applications
- Chronic Myelogenous Leukemia (CML)
- Myelodysplastic Syndrome (MDS)
- Acute Lymphoblastic Leukemia (ALL)

Normal Cell:
Two green (2G) and two orange (2O) signals.

Aberrant Cell (typical result):
One green (1G), three orange (3O) signals, indicating the presence of i(17q).

Aberrant Cell (typical results):
Two green (2G), three orange (3O) signals, indicating a gain of 17q.

Aberrant Cells (typical results):
One green (1G) and two orange (2O) signals, indicating a deletion of the TP53 locus.
- Fioretos et al (1999) Blood 94:225-232
- Barbouti et al (2004) Am J Hum Genet 74:1-10
- Carvalho et al (2008) Genome Res 18:1724-1732
Certificate of Analysis (CoA)
or go to CoA Database