XL TCRA/D
Break Apart Probe
- Order Number
- D-5106-100-OG
- Package Size
- 100 µl (10 Tests)
- Chromosome
- 1414
- Regulatory Status
- IVDD
IVDR Certification
MetaSystems Probes has already certified a wide range of FISH probes, according to IVDR.
This product remains IVDD-certified until further notice.
Discover all IVDR-certified products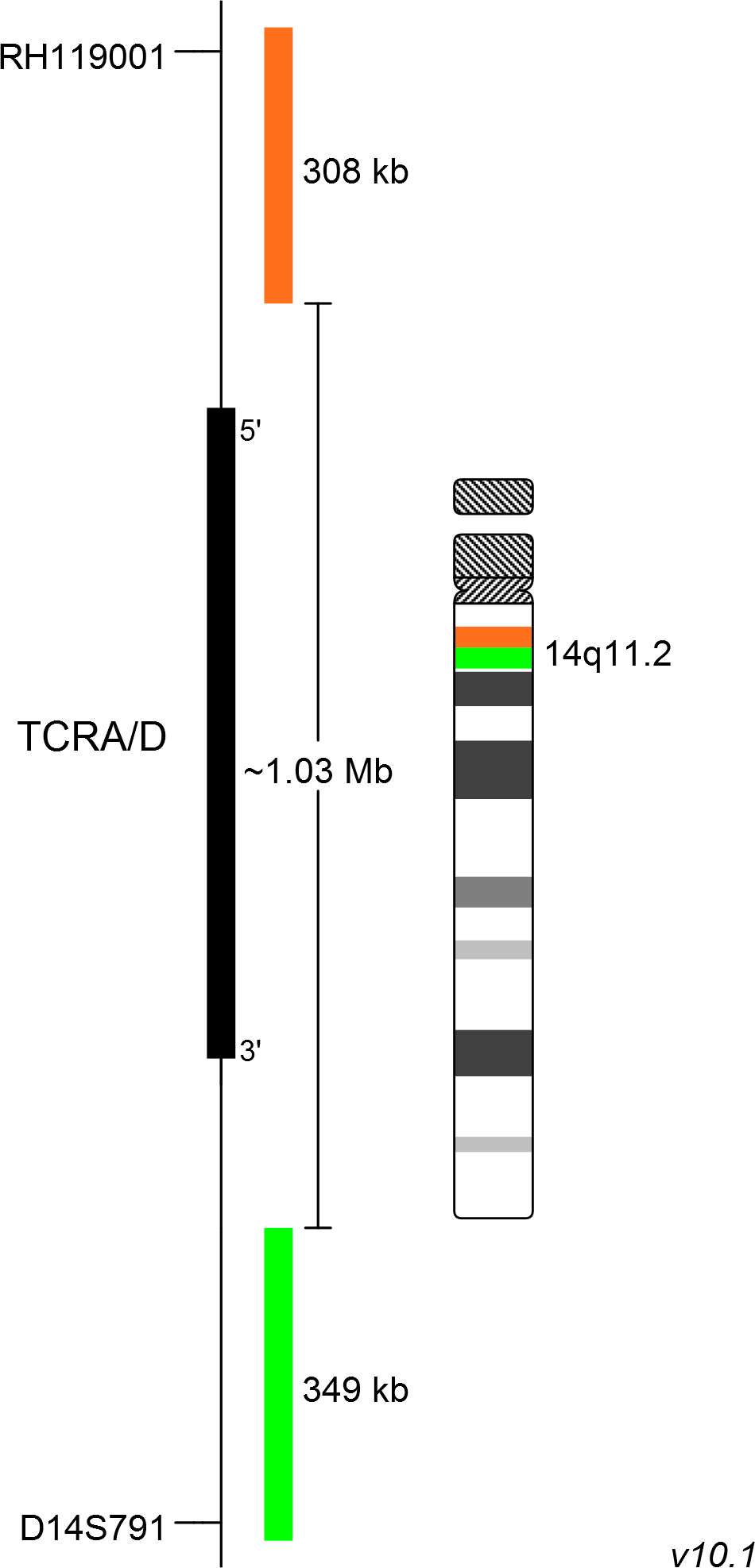
XL TCRA/D consists of an orange-labeled probe hybridizing proximal to the TCRA/D gene region at 14q11.2 and a green-labeled probe hybridizing distal to TCRA/D gene region at 14q11.2.
Probe maps for selected products have been updated. These updates ensure a consistent presentation of all gaps larger than 10 kb including adjustments to markers, genes, and related elements. This update does not affect the device characteristics or product composition. Please refer to the list to find out which products now include updated probe maps.
Probe map details are based on UCSC Genome Browser GRCh37/hg19, with map components not to scale.
Chromosomal aberrations with breakpoints in T-cell receptor (TCR) gene loci are recurrent in several T-cell malignancies. The chromosomal alterations juxtapose oncogenes next to TCR regulatory sequences leading to deregulated expression of those oncogenes.
T-cell prolymphocytic leukemia (T-PLL) harbors frequent alterations of the TCRA/D locus, usually caused by an inv(14)(q11q32). By molecular cytogenetic studies, the incidence of TCRA/D rearrangements is about 24% of all T-cell acute lymphoblastic leukemia (T-ALL) cases.
Clinical Applications
- Acute Lymphoblastic Leukemia (ALL)
- Non-Hodgkin Lymphomas (NHL)

Normal Cell:
Two green-orange (2GO) fusion signals representing the two normal TCRA/D loci.

Aberrant Cell (typical results):
One green (1G), one orange (1O), and one green-orange (1GO) fusion signal, indicating a chromosome break in the TCRA/D locus.
- Gesk et al (2003) Leukemia 17:738-745
- Leich et al (2007) J Pathol 213:99-105
- Feldman et al (2009) Am J Clin Pathol 130:178-185
Certificate of Analysis (CoA)
or go to CoA Database

