XL SS18 BA
Break Apart Probe
- Order Number
- D-6033-100-OG
- Package Size
- 100 µl (10 Tests)
- Chromosome
- 1818
- Regulatory Status
- IVDD
IVDR Certification
MetaSystems Probes has already certified a wide range of FISH probes, according to IVDR.
This product remains IVDD-certified until further notice.
Discover all IVDR-certified products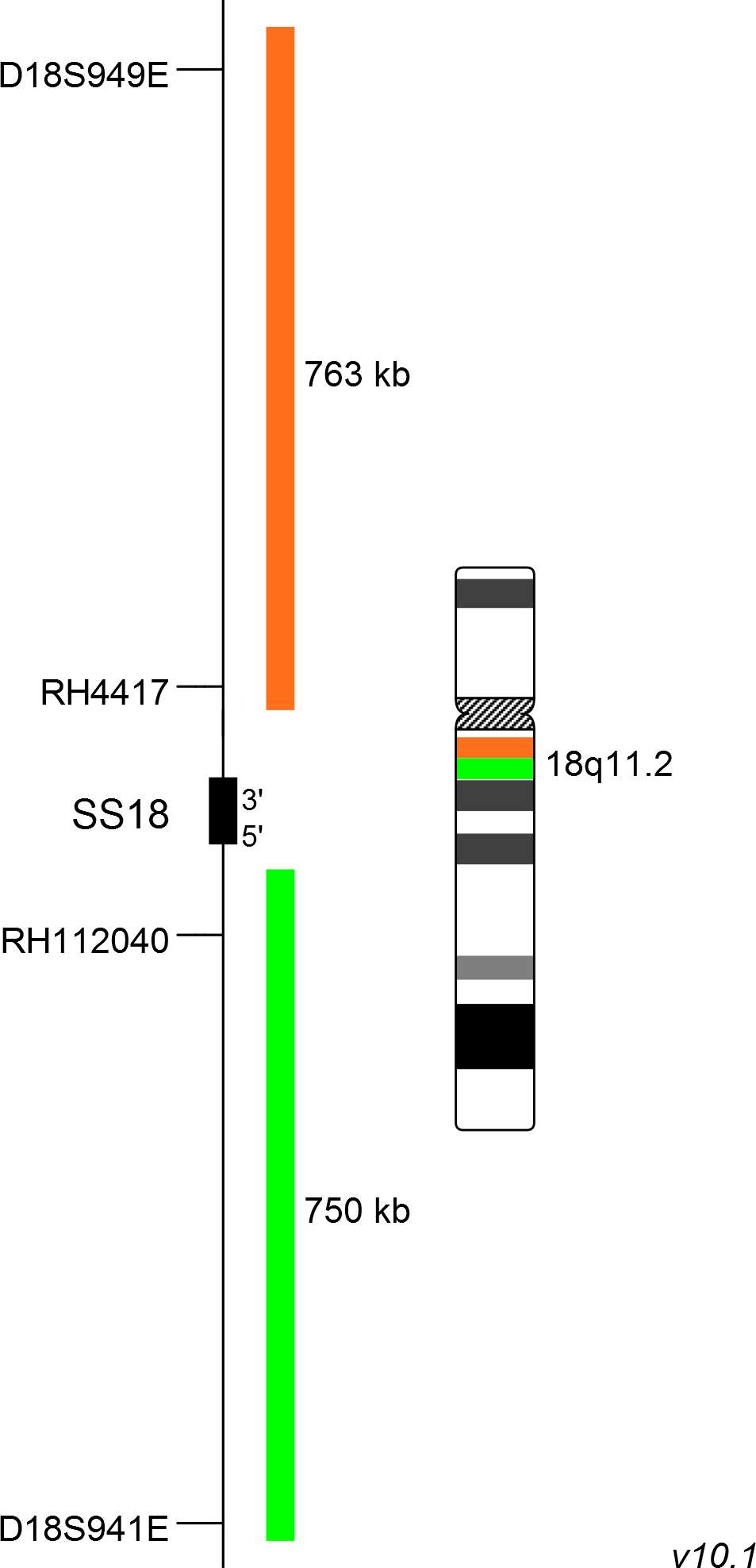
XL SS18 BA consists of an orange-labeled probe hybridizing proximal to the SS18 gene region at 18q11.2 and a green-labeled probe hybridizing distal to the SS18 gene region at 18q11.2.
Probe maps for selected products have been updated. These updates ensure a consistent presentation of all gaps larger than 10 kb including adjustments to markers, genes, and related elements. This update does not affect the device characteristics or product composition. Please refer to the list to find out which products now include updated probe maps.
Probe map details are based on UCSC Genome Browser GRCh37/hg19, with map components not to scale.
Synovial sarcoma is a highly aggressive and relatively rare soft tissue sarcoma. It often develops in the limbs of adolescents and young adults and comprises about 10-20% of soft tissue sarcomas in this population. The disease is characterized by the balanced translocation t(X;18) resulting in an in-frame fusion of ´synovial sarcoma translocation, chromosome 18´ (SS18) with members of the ´synovial sarcoma, X breakpoint´ family (SSX), located on the X chromosome. The chimeric fusion protein consists of almost the complete SS18 protein, only eight amino acids are missing, with the carboxy terminus of SSX1 or SSX2 and in rare cases SSX4. Several studies have shown that the SS18-SSX chimeric protein is interacting with the ATP-dependent chromatin remodeling complex SWI/SNF, interfering with its proper function. The absence of the der(18) in some cases suggests that it is not mandatory for the maintenance of the tumor. Generally, synovial sarcoma shows a low genetic complexity, about 50% of the cases harbor t(X;18) as a sole aberration. Since cytogenetic analysis of solid tumors is challenging, FISH provides an alternative method for the characterization of the status of SS18-SSX rearrangements.
Clinical Applications
- Solid Tumors (Solid Tumors)

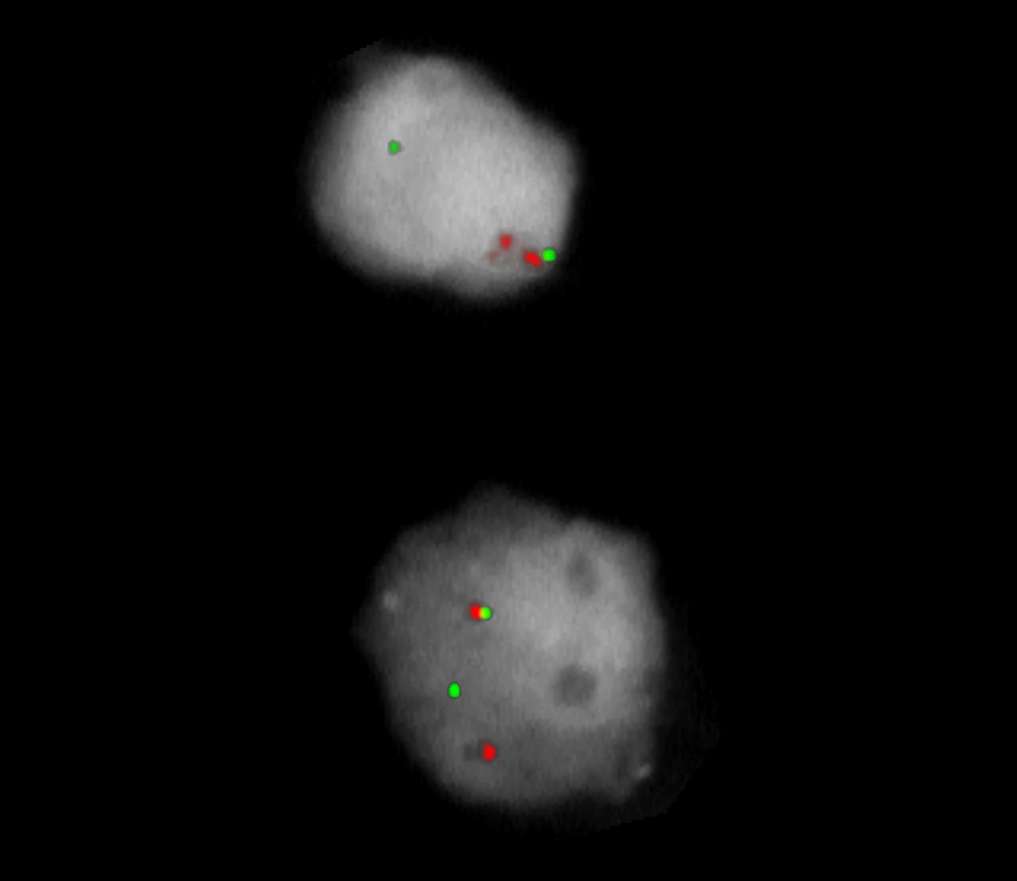
Normal Cell:
Two green-orange colocalization/fusion signals (2GO).

Aberrant Cell (typical results):
One green-orange colocalization/fusion signal (1GO), one separate green (1G) and orange (1O) signal each resulting from a chromosome break in the relevant locus.
- Shipley et al (1996) Am J Pathol 148:559-567
- Surace et al (2004) Lab Invest 84:1185-1192
- Nielsen et al (2015) Cancer Discov 5:124-134
Certificate of Analysis (CoA)
or go to CoA Database