XL FUS BA
Break Apart Probe
- Order Number
- D-6035-100-OG
- Package Size
- 100 µl (10 Tests)
- Chromosome
- 1616
- Regulatory Status
- IVDD
IVDR Certification
MetaSystems Probes has already certified a wide range of FISH probes, according to IVDR.
This product remains IVDD-certified until further notice.
Discover all IVDR-certified products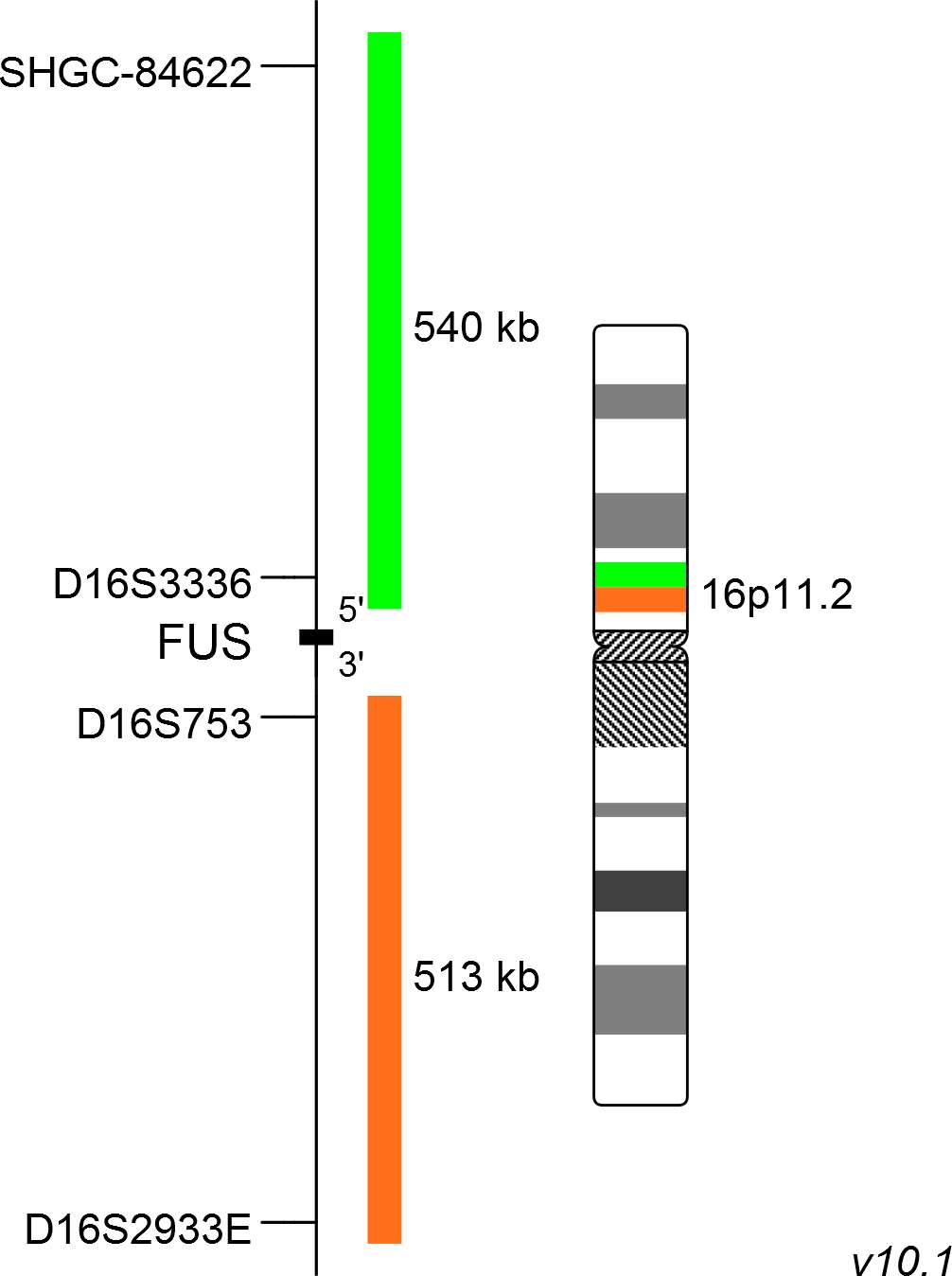
XL FUS BA consists of an orange-labeled probe hybridizing proximal to the FUS gene region at 16p11.2 and a green-labeled probe hybridizing distal to the FUS gene region at 16p11.2.
Probe maps for selected products have been updated. These updates ensure a consistent presentation of all gaps larger than 10 kb including adjustments to markers, genes, and related elements. This update does not affect the device characteristics or product composition. Please refer to the list to find out which products now include updated probe maps.
Probe map details are based on UCSC Genome Browser GRCh37/hg19, with map components not to scale.
Myxoid liposarcomas (MLS) are accounting for about 30% of liposarcomas and represent approximately 10% of adult soft tissue sarcomas. Patients with MLS showing progression to round-cell morphology have an inferior outcome. The most common aberration in MLS is the translocation t(12;16)(q13;p11) with a frequency of about 95% and to a much lesser extend t(12;22)(q13;q12), in which FUS is not involved. These reciprocal translocations are resulting in the generation of FUS-DDIT3 and EWSR1-DDIT3 fusion genes, respectively. As FUS is also involved in the development of low-grade fibromyxoid sarcoma with the rearrangement t(7;16)(q33;p11), FUS rearrangements are not highly specific for the detection of MLS. Human models of sarcomagenesis suggest that the FUS-DDIT3 fusion gene impedes with adipogenic differentiation of mesenchymal stem cells and thereby contributes to the development of liposarcoma.
Clinical Applications
- Solid Tumors (Solid Tumors)

Normal Cell:
Two green-orange colocalization/fusion signals (2GO).

Aberrant Cell (typical results):
One green-orange colocalization/fusion signal (1GO), one separate green (1G) and orange (1O) signal each resulting from a chromosome break in the relevant locus.
- Knight et al (1995) Cancer Res 55:24-27
- Tanas et al (2009) Adv Anat Pathol 16:383-391
- Rodriguez et al (2013) Stem Cells 31:2061-2072
Certificate of Analysis (CoA)
or go to CoA Database


