XL MALT1 BA
Break Apart Probe
- Order Number
- D-6015-100-OG
- Package Size
- 100 µl (10 Tests)
- Chromosome
- 1818
- Regulatory Status
- IVDD
IVDR Certification
MetaSystems Probes has already certified a wide range of FISH probes, according to IVDR.
This product remains IVDD-certified until further notice.
Discover all IVDR-certified products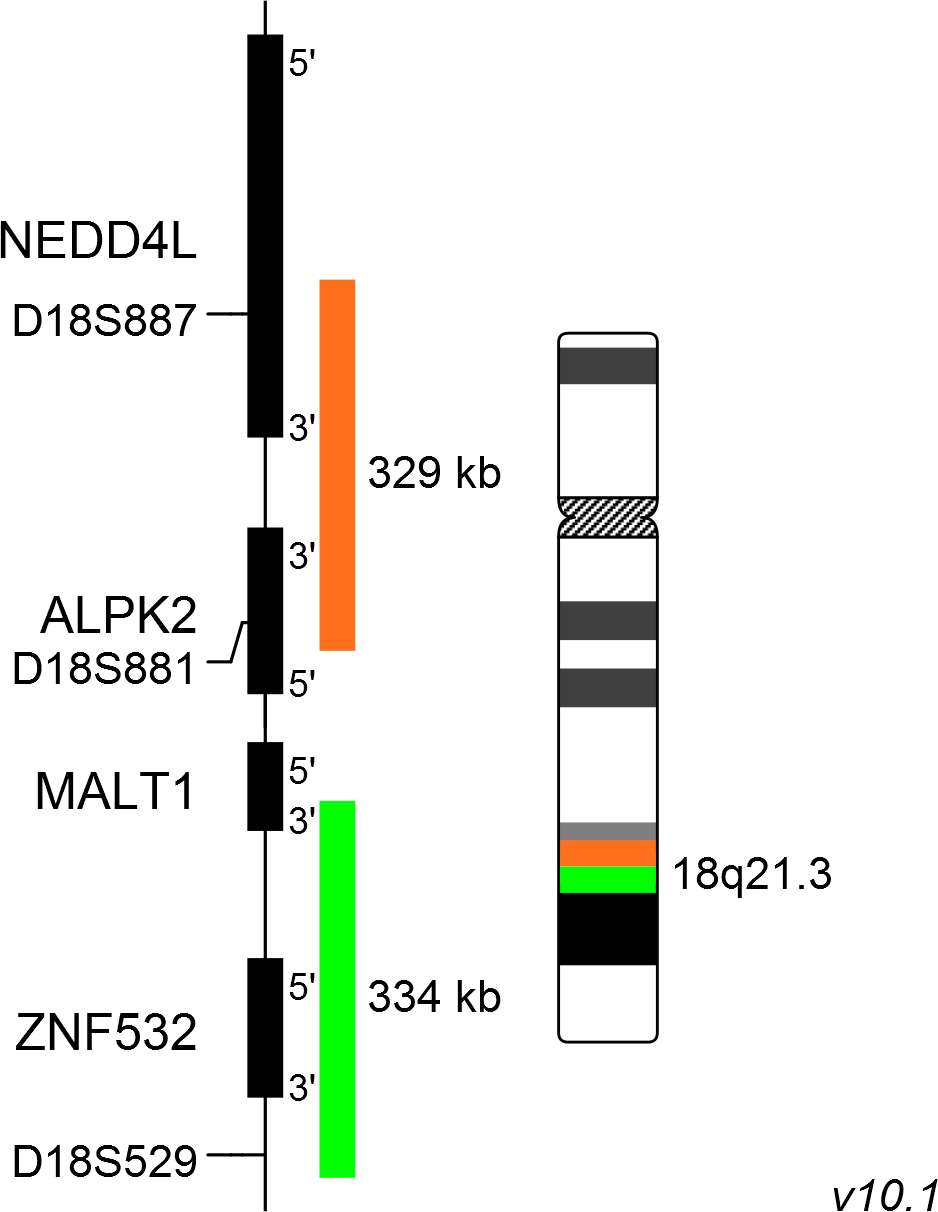
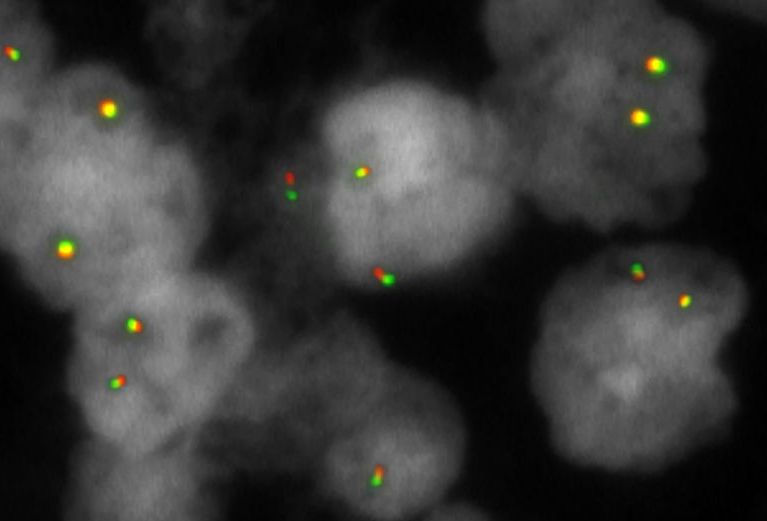
XL MALT1 BA consists of an orange-labeled probe hybridizing proximal to the MALT1 gene region at 18q21.3 and a green-labeled probe hybridizing distal to the MALT1 gene region at 18q21.3.
Probe maps for selected products have been updated. These updates ensure a consistent presentation of all gaps larger than 10 kb including adjustments to markers, genes, and related elements. This update does not affect the device characteristics or product composition. Please refer to the list to find out which products now include updated probe maps.
Probe map details are based on UCSC Genome Browser GRCh37/hg19, with map components not to scale.
The MALT1 gene was identified through its involvement in t(11;18)(q21;q21), seen in 30% of cases of mucosa-associated lymphoid tissue (MALT) lymphoma. The t(11;18)(q21;q21) is restricted to MALT lymphomas and has not been detected in nodal or splenic marginal zone lymphomas, diffuse large B-cell lymphomas, or other non-Hodgkin lymphomas. The second most frequent translocation identified in MALT lymphoma is the t(14;18)(q32;q21) IGH/MALT1. The t(14;18)(q32;q21) IGH/MALT1 is found most often in MALT lymphomas arising at non-gastric sites and is identified in 5-25% of cases arising in the ocular adnexa, lung, salivary gland and skin.
The oncogenic activity of MALT1 is linked to its involvement of the CARMA1-BCL10-MALT1 (CBM) complex in antigen receptor-mediated activation of the transcription factor NF-kB, which controls the expression of numerous anti-apoptotic and proliferation-promoting genes.
Clinical Applications
- Non-Hodgkin Lymphomas (NHL)
- Solid Tumors (Solid Tumors)

Normal Cell:
Two green-orange (2GO) fusion signals representing the two normal MALT loci.

Aberrant Cell (typical results):
One green (1G), one orange (1O), and one green-orange (1GO) fusion signal, indicating a chromosome break in the MALT locus.
- Streubel et al (2003) Blood 101:2335-2339
- Murga Penas et al (2003) Leukemia 17:2225-2229
- Bacon et al (2007) J Clin Pathol 60:361-372
Certificate of Analysis (CoA)
or go to CoA Database